A significant challenge for biological electron microscopists has always been the interpretation of complex structures from two-dimensional images. Scanning electron microscopy (SEM) is normally used for imaging the surface of cells, tissues and whole multicellular organisms. SEM images of surfaces appear to be three-dimensional (3D) but there is no measurable depth information in the image. Improvements in imaging and computing technology have resulted in the development of several SEM techniques that enable high resolution 3D analysis of biological samples.
Backscattered electron image of a wasp. While the image appears 3D, it contains no depth information and cannot be measured.
Stereoscopy and photogrammetry are techniques that use images of the same area of sample captured at different angles, recombining them into a 3D model of the surface. Surface profiles and depth measurements are possible on the modelled data. These techniques are limited in their scope but useful for samples that cannot be damaged, such as museum specimens, or where analysis of the surface topography is important.
Red-cyan stereoscopic image of a bee. Using stereoscopic glasses depth information can be obtained from the image. Images can be processed digitally to provide a 3D model for detailed measurement and analysis.
Detailed ultrastructural information, traditionally only possible using transmission electron microscopy (TEM), can now be collected using high resolution field emission SEMs on frozen and resin embedded samples with a either a flat surface (trimmed with a glass or diamond knife) or cut into sections. Backscattered electrons provide atomic contrast in the sample, resulting in inverted TEM-like images. There are three main techniques for collecting 3D ultrastructural data in an SEM; Serial section array tomography, serial block face SEM (SBFSEM) and focused ion beam SEM (FIBSEM).
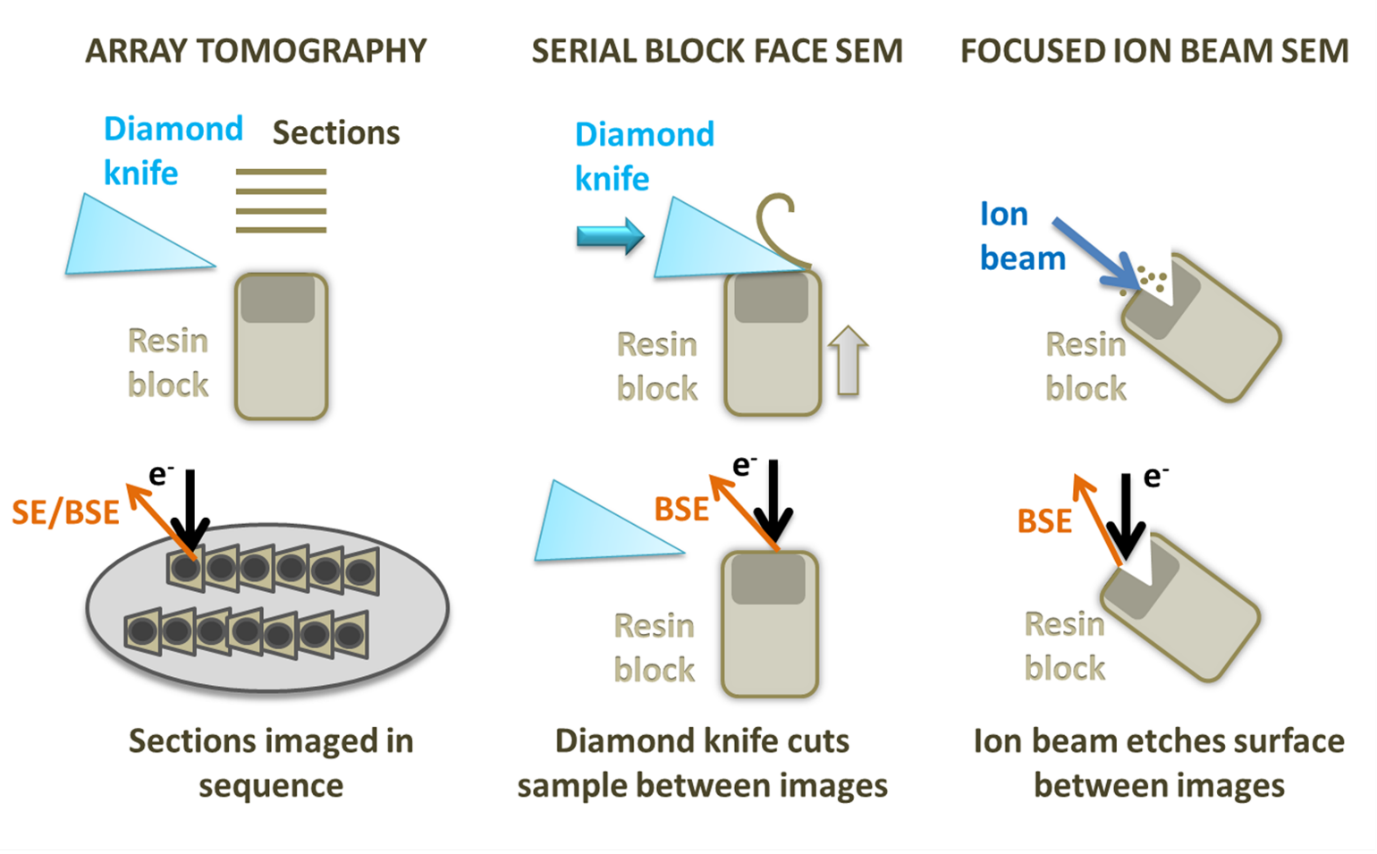
Diagram showing array tomography, serial block face Sem and focused ion beam SEM.
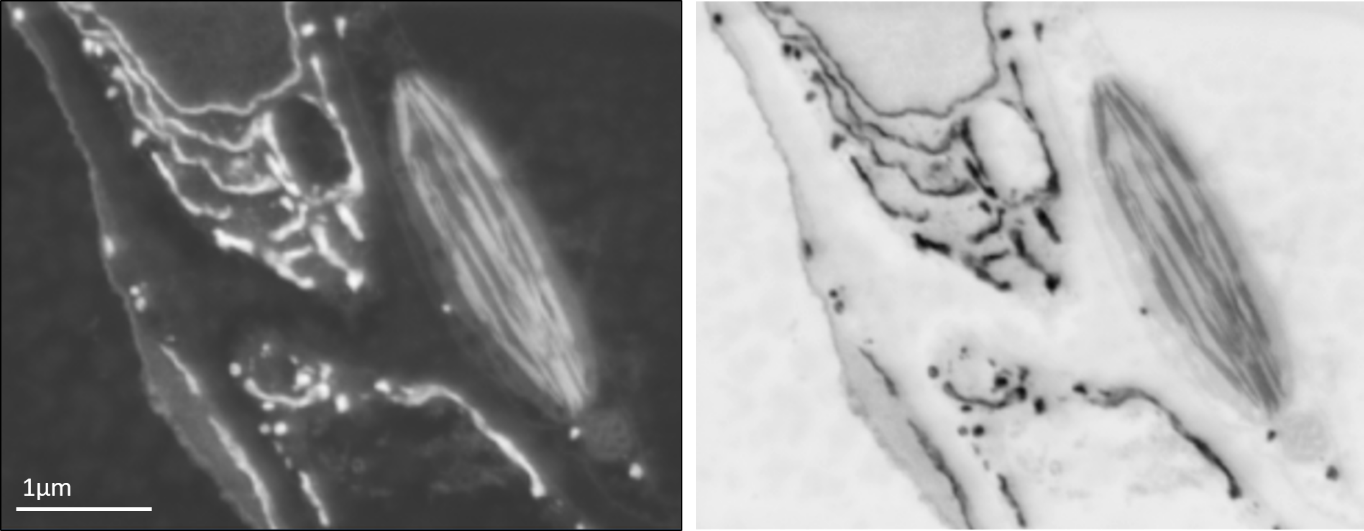
Backscattered electron SEM image of leaf tissue showing endoplasmic reticulum and a chloroplast (left) and the same image inverted for an image that appears similar to the type of data obtained using a TEM (right).
FIBSEM
FIBSEM utilises an ion beam to ablate the sample, creating a trench with a flat surface on one side that can be used for imaging with an electron beam. Once an image of the surface is collected, the ion beam is used to mill away a thin layer (3-5nm) from the imaging region. A second image is captured, and the process is repeated, building up a stack of images. FIBSEM is also able to mill the sample to create a thin lamella that can be used as a TEM section, which is particularly useful for cryo applications. FIBSEM provides the best resolution in Z of all the 3D SEM techniques and can be used on frozen-hydrated samples and resin embedded room temperature specimens. However, it also has the smallest field of view and lowest volume capacity of these methods and has the additional disadvantage of destroying the region of the sample being imaged.
SBFSEM
SBFSEM uses a diamond knife mounted onto a microtome within the SEM specimen chamber to slice the surface of the specimen. The cut face of the sample is imaged before the knife is used to trim another section away. Slices as thin as 15nm can be removed from very stable samples, but more often slices range from 20 to 100nm. This means that the X and y resolution is usually better than the z resolution. On some samples and microscopes, it is possible to recover some extra z resolution using variable voltages for the electron beam. As with FIBSEM, SBFSEM is a destructive method and it is not possible to revisit sections later. SBFSEM is also restricted to room temperature samples with a lot of contrast. SBFSEM can image a relatively large volume of tissue (hundreds of microns) and a large surface area, avoiding section specific artefacts such as compression and wrinkling. The resin block can be prone to beam damage and charging, requiring measures such as using a gas injection system or variable pressure in the microscope or very low accelerating voltages, which reduces contrast. SBFSEM is the fastest method for set up and data collection over a specified volume of all the techniques.
Array tomography
Array tomography involves producing serial sections on an ultramicrotome and mounting them onto a stub or wafer before inserting them into the microscope. Each section is imaged in sequence and the same region can be revisited and imaged multiple times at different magnifications and resolution. Stains and immunolabels can be applied to the sample after embedding, and after imaging once regions of interest have been located, something that cannot currently be done with SBFSEM. Array tomography does not have a limit on the volume of tissue that can be imaged and is the most flexible system. However, the z resolution is restricted by the thickness of the sections (30nm at best) and it can be prone to wrinkles, compression and other section-specific artefacts.
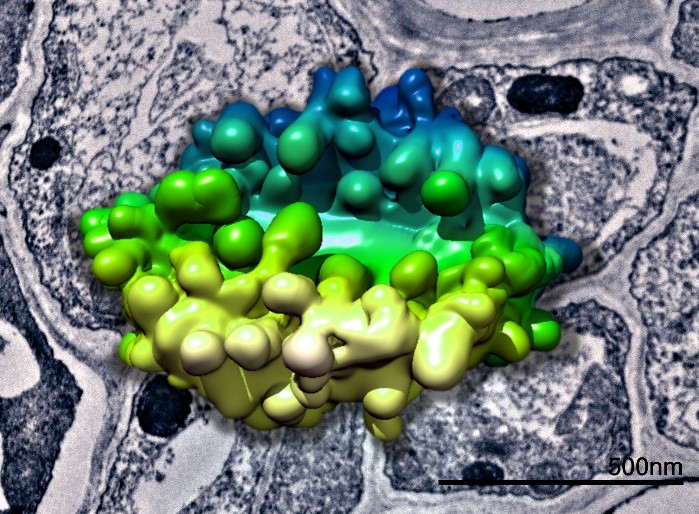
3D model of a plant Golgi body generated from SBFSEM data superimposed above a TEM image of plant tissue. Data collected in collaboration with Chris Hawes.
All the 3D SEM techniques produce large volumes of data that require significant processing and analysis time. However, the advantages of being able to visualise biological ultrastructure in 3D cannot be emphasized enough, avoiding misinterpretation of key structural data, which makes investment in time and equipment worth the effort. It is an exciting time to be involved in this rapidly developing field.
To learn more about not only 3D SEM for the visualisation and analysis of biological samples, download our recent whitepaper highlighting the benefits of EDS for life sciences.