2nd June 2021 | Author: Dr Louise Hughes
Key techniques in advanced biological SEM
Biological SEM has come into its own over the past decade, moving from a relatively niche application looking solely at surface structure to being able to address internal tissue and cell ultrastructure in three-dimensions at resolutions approaching that of TEM. There are several techniques of particular interest to biologists.
STEM in SEM
Contrast in biological EM is an ongoing challenge. There are numerous methods to address this, including modifying sample preparation to improve contrast, using compositional information (see multi-colour SEM below) and by optimising microscope operating conditions. Lowering the accelerating voltage of a TEM significantly improves the contrast of a sample but cannot reach the low accelerating voltages available with SEMs (<1-30kV). Dedicated STEM holders and STEM detectors in an SEM replicate the detection of electrons transmitted through a thin section, resulting in the ability to image low contrast samples, reduce sample contamination, reduce specimen charging, and potentially reduce damage to specimens. There is a trade off in that STEM in SEM does not have the same resolution that can be achieved in a TEM, but it also expands the capabilities of SEM for slightly lower resolution applications.
Block-face imaging
Ultrastructural information about a sample can also be obtained from the flat surface of a specimen embedded in resin. Backscattered electron images can be obtained from the cut or polished surface of the resin block and reveal ultrastructural information about the sample. The advantage of block face imaging over TEM or STEM in SEM is the large surface area that can be imaged without interruption by supporting materials such as grid bars. Very low accelerating voltages with long beam dwell times can achieve high resolution images. Beam sensitivity and charging issues mean that imaging conditions are critical to achieve the best imaging results on these types of sample.
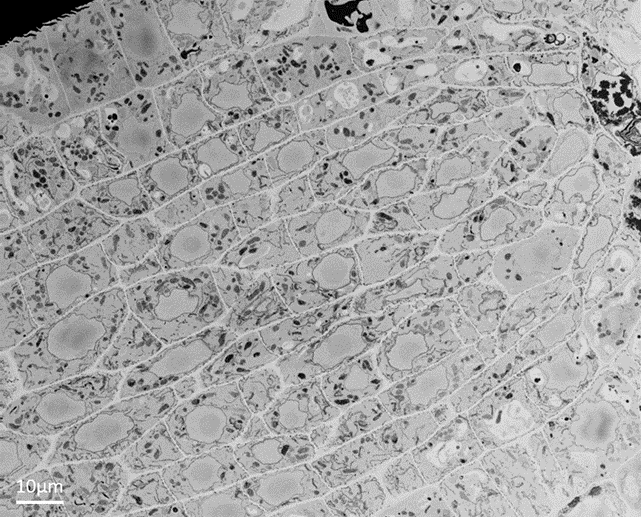
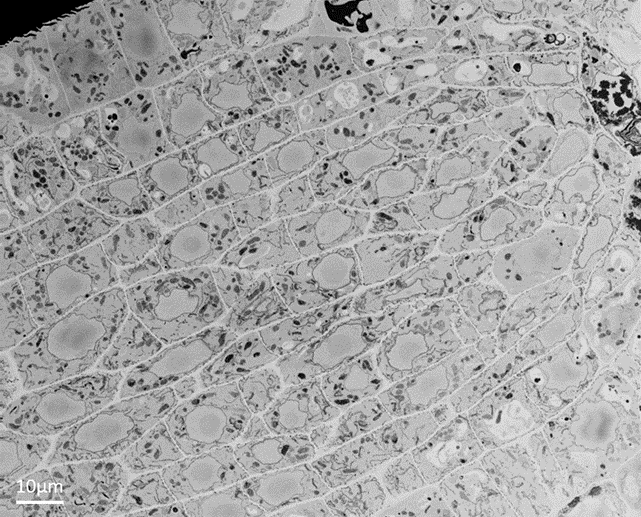
Figure 1. Block-face image of an Arabidopsis root tip imaged with backscattered electrons and showing cell ultrastructure.
Volume SEM
Volume microscopy can be done in a TEM (serial section TEM and electron tomography), but only for a relatively small volume of a sample, such as part of a cell or organelle. For volumes requiring data across whole cells, large areas of tissue, or whole organisms at ultrastructural resolutions, volume SEM offers a range of techniques. Details of volume SEM techniques can be found in a previous blog post. The three most commonly used techniques involve slicing the sample and capturing images of either the slices or the remaining specimen block (serial block-face imaging).
- Focused ion beam scanning electron microscopy (FIBSEM) uses a dual beam microscope. An electron beam captures images of the surface of a sample. The ion beam ablates a thin layer away from the surface in between the acquisition of electron images.
- Serial block face scanning electron microscopy (SBFSEM) involves removing the surface layer of the sample using a diamond knife between images.
- Array tomography (or serial section SEM) involves creating a series of slices or sections outside of the microscope, preserving them on a substrate and sequentially imaging each slice.
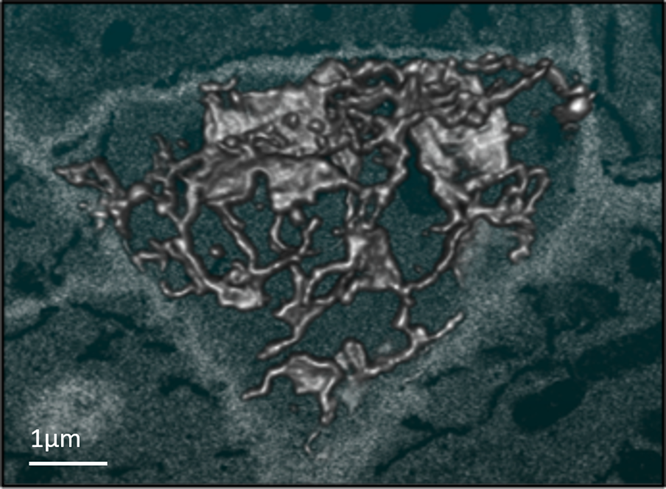
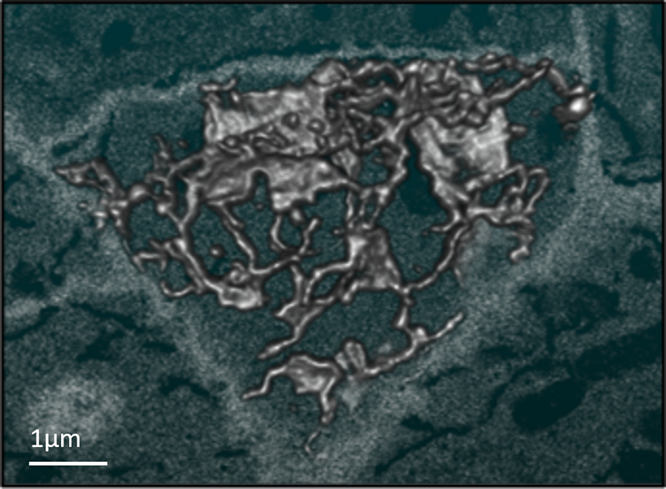
Figure 2. SBFSEM data showing the 3D reconstruction of endoplasmic reticulum at the edge of a root cell. Data captured from the same specimen shown in figure 1.
Each of the volume SEM techniques provides data that facilitates the accurate interpretation and modelling of complex 3D ultrastructure that would be very difficult to analyse using traditional 2D electron images.
Cryo-SEM
Cryo-SEM uses specially cooled stages and sample transfer equipment to image samples as close to their natural state as possible, samples that are fully hydrated and frozen. A large challenge with all biological electron microscopy is the extensive sample preparation required to preserve a specimen exposed to the harsh electron microscope environment of high vacuums and damaging electron beams. Rapidly freezing a sample to avoid ice-crystal formation, and thus vitrify the water content, provides the fastest preservation possible and creates the fewest artefacts in the specimen. Cryo-SEM can be used in combination with other techniques, such as correlative light and electron microscopy (CLEM) or FIBSEM to either image the frozen sample in 3D or to create slices of thin, frozen samples for high resolution TEM.
Multi-colour SEM
It is possible to image structural information using electron signals and combine this with compositional information about biological specimens. This compositional data can be obtained using energy dispersive x-ray spectrometry (EDS), electron energy loss spectroscopy (EELS) and cathodoluminescence, and can be applied to electron data through the application of colour. These techniques can be used to identify elements found in the sample, map their distribution, and measure their relative quantity. More information about EDS for biological research is on our application pages.
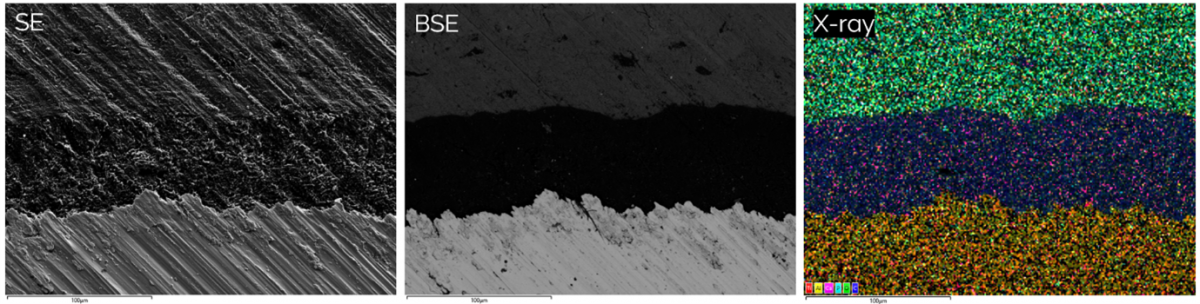

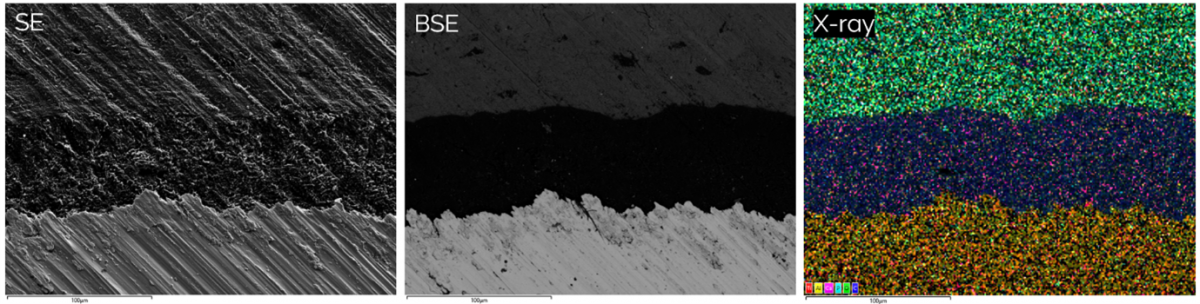
Figure 3 showing the differences between topographical secondary electron (SE) images, backscattered electron images (BSE) showing differences in density and x-ray signals revealing the distribution of elements in a bone implant (sample courtesy of Zhidao Xia, Swansea University).
The range of SEM techniques available to biologists means that SEM is now a key tool in any biological imaging facility. If you would like to learn about advanced biological SEM in more detail, register for our upcoming webinar, where I will be talking about these topics as well as how to make the most out of your microscope by optimising imaging conditions and sample preparation techniques.